Pandaral·lel
A simple and efficient tool to parallelize Pandas operations on all available CPUs.
pandarallel is a simple and efficient tool to parallelize Pandas operations on all
available CPUs.
With a one line code change, it allows any Pandas user to take advandage of his
multi-core computer, while pandas uses only one core.
pandarallel also offers nice progress bars (available on Notebook and terminal) to
get an rough idea of the remaining amount of computation to be done.
| Without parallelization |  |
|---|---|
| With parallelization |  |
Features
pandarallel currently implements the following pandas APIs:
| Without parallelization | With parallelization |
|---|---|
df.apply(func) |
df.parallel_apply(func) |
df.applymap(func) |
df.parallel_applymap(func) |
df.groupby(args).apply(func) |
df.groupby(args).parallel_apply(func) |
df.groupby(args1).col_name.rolling(args2).apply(func) |
df.groupby(args1).col_name.rolling(args2).parallel_apply(func) |
df.groupby(args1).col_name.expanding(args2).apply(func) |
df.groupby(args1).col_name.expanding(args2).parallel_apply(func) |
series.map(func) |
series.parallel_map(func) |
series.apply(func) |
series.parallel_apply(func) |
series.rolling(args).apply(func) |
series.rolling(args).parallel_apply(func) |
Requirements
On Linux & macOS, no special requirement.
On Windows, because of the multiprocessing system (spawn), the function you send to
pandarallel must be self contained, and should not depend on external resources.
Example:
✅ Valid on Mac and Linux - ❌ Forbidden On Windows
import math
def func(x):
# Here, `math` is defined outside `func`. `func` is not self contained.
return math.sin(x.a**2) + math.sin(x.b**2)
✅ Valid everywhere
def func(x):
# Here, `math` is defined inside `func`. `func` is self contained.
import math
return math.sin(x.a**2) + math.sin(x.b**2)
Warning
Parallelization has a cost (instantiating new processes, sending data via shared memory, ...), so parallelization is efficient only if the amount of computation to parallelize is high enough. For very little amount of data, using parallelization is not always worth it.
Warning
Displaying progress bars has a cost and may slighly increase computation time.
Examples
An example of each available pandas API is available:
- For Mac & Linux
- For Windows
Benchmark
For some examples, here is the comparative benchmark with and without using Pandaral·lel.
Computer used for this benchmark:
- OS: Linux Ubuntu 16.04
- Hardware: Intel Core i7 @ 3.40 GHz - 4 cores
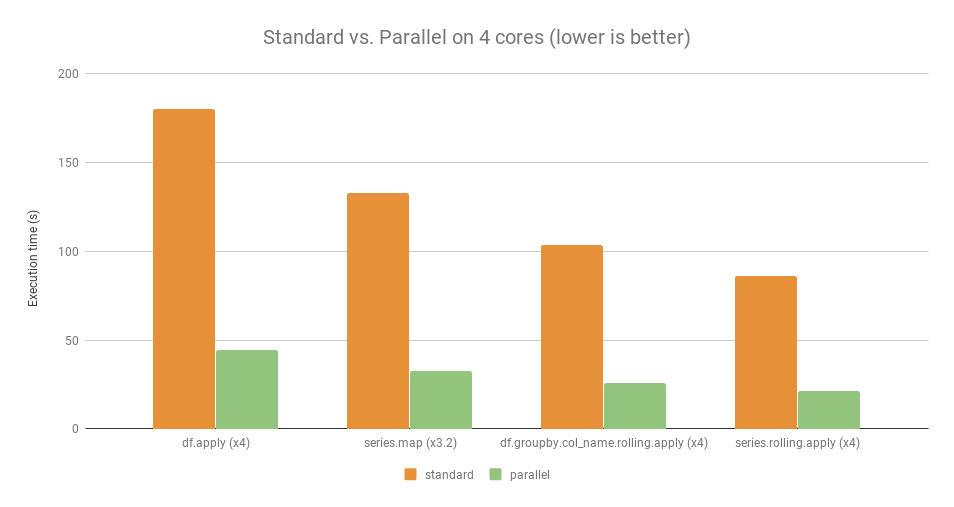
For those given examples, parallel operations run approximately 4x faster than the standard operations (except for series.map which runs only 3.2x faster).
When should I user pandas, pandarallel or pyspark?
According to pandas documentation:
pandasis a fast, powerful, flexible and easy to use open source data analysis and manipulation tool,built on top of the Python programming language.
The main pandas drawback is the fact it uses only one core of your computer, even if
multiple cores are available.
pandarallel gets around this limitation by using all cores of your computer.
But, in return, pandarallel need twice the memory that standard pandas operation
would normally use.
==> pandarallel should NOT be used if your data cannot fit into memory with
pandas itself. In such a case, spark (and its python layer pyspark)
will be suitable.
The main drawback of spark is that spark APIs are less convenient to user than
pandas APIs (even if this is going better) and also you need a JVM (Java Virtual
Machine) on your computer.
However, with spark you can:
- Handle data much bigger than your memory
- Using a
sparkcluster, distribute your computation over multiple nodes.

